Carmelo Rizzo
Professor of Chemistry
Chemistry and Biology of DNA Damage.
Covalent modification of DNA by electrophiles is generally accepted as the initial event in chemical carcinogenesis. If these modifications are not repaired, they can compromise the fidelity of DNA replication leading to mutations and possibly cancer. To properly study the biological processing of pre-mutagenic DNA lesions, oligonucleotides containing structurally defined carcinogen adducts are required. Our laboratory develops synthetic strategies for the site-specific incorporation of nucleotides that are chemically modified by carcinogens into DNA. Once synthesized, the structure, replication and repair of the carcinogen-modified oligonucleotides are examined. Many of these studies are preformed in collaboration with other laboratories on Vanderbilt's campus and elsewhere.
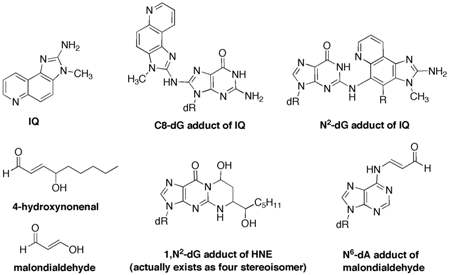
One specific project includes the preparation of the C8-deoxyguanosine adduct of 2-amino-3-methylimidazo[4,5-f]quinoline (IQ). IQ is a member of a family of highly mutagenic heterocyclic amines found in cooked meats. We found that the C8-IQ adduct adopts different conformations depending on the sequence of the adducted oligonucleotide and we have hypothesized that this sequence-dependent conformation plays an important role in the biological processing of the lesion. This hypothesis will be tested using in vitro and in vivo systems. The adduction of IQ with DNA also gives a minor N2-adduct of deoxyguanosine, which has not been extensively studied. We have recently completed a synthesis of the N2-adduct and have incorporated it into oligonucleotides.
A second major project in our lab involves DNA adducts derived from endogneous sources such as lipid peroxidation. Examples of such reactive electrophiles include alpha,beta-unsaturated aldehydes (acrolein, crotonaldehyde, and 4-hydroxynonenal), 2,3-epoxyaldehydes and dicarbonyl species (malondialdehyde and 4-oxo-2-nonenal). Although these compounds are chemically simple, they react with DNA in a complex and diverse manner. We recently demonstrated that alpha,beta-unsaturated aldehydes can form inter- and intra-strand DNA crosslinks, which are a very severe but largely unstudied form of DNA damage. The crosslinking chemistry is highly dependent on the stereochemistry of the DNA adduct. In collaboration with other laboratories, we are studying the mechanism of the DNA crosslinking reaction and the biological processing of the DNA crosslinks.
Modern mass spectrometry and NMR are integral parts to all of our studies on DNA damage. We have convenient and open access to superb analytical facilities on Vanderbilt campus. Our laboratory is affiliated with the Vanderbilt Center in Molecular Toxicology, Vanderbilt Institute of Chemical Biology, and the Vanderbilt-Ingram Cancer Center .
Specializations
VICB
Organic Chemistry
Chemical Biology
Bioorganic Chemistry
Representative Publications
Minko, I.G., Rizzo, C.J., Lloyd, R.S. (2017) Mutagenic Potential of Nitrogen Mustard Formamidopyrimidine DNA Adduct, J. Biol. Chem. 292, 18790-18799. PMCID: PMC5704464
Sha, Y., Minko, I.G., Malik, C.K., Rizzo. C.J., Lloyd, R.S. (2017) Error-prone replication bypass of the imidazole ring-opened formamidopyrimidine deoxyguanosine adduct, Environ. Mol. Mutagen. 58, 182-189. (feature article). PMCID: PMC5476229
Bose, A., Millsap, A..D., DeLeon A., Rizzo, C.J., Basu, A .K. (2016) Translesion Synthesis of the N2-2′-Deoxyguanosine Adduct of the Dietary Mutagen IQ in Human Cells: Error-Free Replication by DNA Polymerase κ and Mutagenic Bypass by DNA Polymerases η, ζ, and Rev1, Chem. Res. Toxicol. 29, 1549-1559. (cover article) PMCID: PMC5031085
Minko, I. G., Jacobs, A. C., Donley, N., Gruppi,F., Harris, T. M., Rizzo,C. J., McCullough, A. K,. Lloyd, R. S. (2016) Catalysts of DNA strand cleavage at apurinic/apyrimidinic sites, Sci. Rep.(Nature), 6, 28894; doi: 10.1038/srep28894. PMCID: PMC4929455
Thiaville, J. J., Kellner, S. M., Yuan, Y., Hutinet, G., Thiaville, P. C., Jumpathong, W., Mohapatra, S., Brochier-Armanet, C., Letarov, A. V., Hillebrand, R., Malik, C. K., Rizzo, C. J. Dedon, P. C., de Crécy-Lagard, V. (2016) Novel genomic island modifies DNA with 7-deazaguanine derivatives, Proc. Natl. Acad. Sci. U.S.A., 113, E1452–E1459. PMCID: PMC4801273
Gruppi, F., Hejzai, L., Christov, P. P., Krishnamachari, S., Turesky, R. J., Rizzo, C. J. (2015) Characterization of nitrogen mustard formamidopyrimidine adduct formation of bis-(2-chloroethyl)ethylamine with calf thymus DNA and human mammary cancer cell line, Chem. Res. Toxicol. 28, 1850-1860. PMCID: PMC4579055
Bose, A., Pande, P., Jasti, V. P., Millsap, A. D., Hawkins, E. K., Rizzo, C. J., Basu, A. K. (2015) DNA polymerases κ and ζ cooperatively perform mutagenic translesion synthesis of the C8-2’-deoxyguanosine adduct of the dietary mutagen IQ in human cells, Nucleic Acids Res. 43, 8340-8351. PMID: 26220181
Stavros, K. M., Hawkins, E. K., Rizzo, C. J., Stone, M. P. (2015) Base-displaced intercalated conformation of the 2-amino-3-methylimidazo[4,5-f]quinolone N2-dG DNA adduct positioned at the non-reiterated G1 in the NarI restriction site, Chem. Res. Toxicol. 28, 1455-1468. PMCID: PMC4511292
Patra, A., Banerjee, S, Johnson-Salyard, T., Malik, C., Christov, P., Rizzo, C. J., Stone, M. P., Egli, M. (2015) Structural basis for error-free bypass of the 5-N-methyl-formamidopyrimidine-dG lesion by human DNA polymerase η and sulfolobus solfataricus P2 polymerase IV (Dpo4), J. Am. Chem. Soc. 137, 7011-7014. PMCID: PMC45452697
Zdżalik, D., Domańska, A., Prorok, P., Kosicki, K., van den Born, E., Falnes, P. Ø., Rizzo, C. J., Guengerich, F. P., Tudek, B. (2015) Differential repair of etheno-DNA adducts by bacterial and human AlkB proteins, DNA Repair, 30, 1-10. PMCID: PMC4451939
Chang, S.-c., Fedeles, B. I., Wu, J., Delaney, J. C., Li, D., Zhao, L., Christov, P. P., Yau, E., Singh, V., Jost, M., Drennan, C. L., Marnett, L. J., Rizzo, C. J., Levine, S. S., Guengerich, F. P., Essigmann, J. M. (2015) Next-generation sequencing reveals the biological significance of the N2,3-ethenoguanine lesion in vivo, Nucleic Acids Res. 43, 5489-5500. PMCID: PMC44776462
Gruppi F, Johnson Salyard TL, Rizzo CJ. Synthesis of G-N(2)-(CH2)3-N(2)-G Trimethylene DNA Interstrand Cross-Links. Current Protocols Nucleic Acid Chemistry. 2014, 56:5.14.1-5.14.15.
Christov PP, Son KJ, Rizzo CJ. Synthesis and characterization of oligonucleotides containing a nitrogen mustard formamidopyrimidine monoadduct of deoxyguanosine. Chemical Research in Toxicology. 2014, 27(9):1610-8.
Petrova KV, Millsap AD, Stec DF, Rizzo CJ. Characterization of the deoxyguanosine-lysine cross-link of methylglyoxal. Chemical Research in Toxicology. 2014, 27(6):1019-29.
Stavros KM, Hawkins EK, Rizzo CJ, Stone MP. Base-displaced intercalation of the 2-amino-3-methylimidazo[4,5-f]quinolone N2-dG adduct in the NarI DNA recognition sequence. Nucleic Acids Research. 2014, 42 (5): 3450-63.
Maddukuri L, Shuck SC, Eoff RL, Zhao LL, Rizzo CJ, Guengerich FP, Marnett LJ. Replication, Repair, and Translesion Polymerase Bypass of N-6-Oxopropenyl-2 '-deoxyadenosine. Biochemistry. 2013, 52 (48): 8766-8776.
Zhao LL, Pence MG, Christov PP, Wawrzak Z, Choi JY, Rizzo CJ, Egli M, Guengerich FP. Basis of Miscoding of the DNA Adduct N-2,3-Ethenoguanine by Human Y-family DNA Polymerases. Journal of Biological Chemistry. 2013, 287 (42): 35516-35526.
Christov PP, Yamanaka K, Choi JY, Takata KI, Wood RD, Guengerich FP, Lloyd RS, Rizzo CJ. Replication of the 2,6-Diamino-4-hydroxy-N-5-(methyl)-formamidopyrimidine (MeFapy-dGuo) Adduct by Eukaryotic DNA Polymerases. Chemical Research in Toxicology. 2012, 25 (8): 1652-1661.
Zhao LL, Christov PP, Kozekov ID, Pence MG, Pallan PS, Rizzo CJ, Egli M, Guengerich FP. Replication of N2,3-Ethenoguanine by DNA Polymerases. Angewandte Chemie-International Edition. 2012, 51 (22): 5466-5469.
Banerjee S, Christov PP, Kozekova A, Rizzo CJ, Egli M, Stone MP. Replication Bypass of the trans-4-Hydroxynonenal-Derived (6S,8R,11S)-1,N-2-Deoxyguanosine DNA Adduct by the Sulfolobus solfataricus DNA Polymerase IV. Chemical Research in Toxicology. 2012, 25 (2): 422-435.
Huang H, Wang H, Kozekova A, Rizzo CJ, Stone MP. Formation of a N(2)-dG:N(2)-dG Carbinolamine DNA Cross-link by the trans-4-Hydroxynonenal-Derived (6S,8R,11S) 1,N(2)-dG Adduct. Journal of The American Chemical Society. 2011, 133 (40): 16101-16110.
Huang H, Wang H, Voehler MW, Kozekova A, Rizzo CJ, McCullough AK, Lloyd RS, Stone MP. gamma-Hydroxy-1,N(2)-propano-2 '-deoxyguanosine DNA Adduct Conjugates the N-Terminal Amine of the KWKK Peptide via a Carbinolamine Linkage. Chemical Research in Toxicology. 2011, 24 (7): 1123-1133.
Shanmugam G, Kozekov ID, Guengerich FP, Rizzo CJ, Stone MP. 1,N(2)-Etheno-2 '-deoxyguanosine Adopts the syn Conformation about the Glycosyl Bond When Mismatched with Deoxyadenosine.Chemical Research in Toxicology. 2011, 24 (7): 1071-1079.
Petrova KV, Stec DF, Voehler M, Rizzo CJ, Jeong BS. Synthesis of the four stereoisomers of 2,3-epoxy-4-hydroxynonanal and their reactivity with deoxyguanosine. Organic Biomolecular Chemistry. 2011, 9 (6): 1960-71.
Maddukuri L, Eoff RL, Choi J-Y, Rizzo CJ, Guengerich FP, Marnett LJ. In Vitro Bypass of the Major Malondialdehyde- and Base Propenal-Derived DNA Adduct by Human Y-family DNA Polymerases kappa, iota, and Rev1. Biochemistry. 2010, 49 (38): 8415-8424.
Christov PP, Petrova KV, Shanmugam G, Kozekov ID, Kozekova A, Guengerich FP, Stone MP, Rizzo CJ. Comparison of the in Vitro Replication of the 7-(2-Oxoheptyl)4,N-2-etheno-2 '-deoxyguanosine and 1,N-2-Etheno-2 '-deoxyguanosine Lesions by Sulfolobus solfataricus P2 DNA Polymerase IV (Dpo4). Chemical Research in Toxicology. 2010, 23 (8): 1330-1341.
Huang H, Kozekov ID, Kozekova A, Rizzo CJ, McCullough AK, Lloyd RS, Stone MP. Minor Groove Orientation of the KWKK Peptide Tethered via the N-Terminal Amine to the Acrolein-Derived 1,N-2-gamma-Hydroxypropanodeoxyguanosine Lesion with a Trimethylene Linkage. Biochemistry. 2010, 49 (29): 6155-6164.
Yamanaka K, Minko IG, Takata K, Kolbanovskiy A, Kozekov ID, Wood RD, Rizzo CJ, Lloyd RS. Novel Enzymatic Function of DNA Polymerase v in Translesion DNA Synthesis Past Major Groove DNA-Peptide and DNA-DNA Cross-Links. Chemical Research in Toxicology. 2010, 23 (3): 689-695.